- Read reviews, compare customer ratings, see screenshots, and learn more about AppGenome - create desktop app from any Website! Download AppGenome - create desktop app from any Website! For macOS 10.11 or later and enjoy it on your Mac.
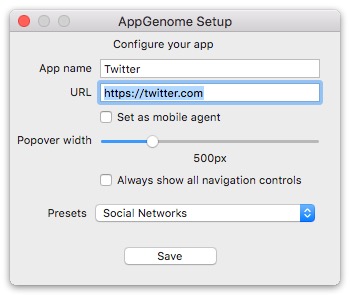
- AppGenome lets you set your own URL, browser agent type (mobile or desktop), and create your own desktop app that will stay in your menubar, so you can forget about all those 'app for' apps. Set your own URL of any web page or web app; Manually set your agent type (browser type): desktop or mobile. This way you choose which version of your website of choice AppGenome to load.
- AppGenome 1.4.1 - lets you set your own URL, browser agent type (mobile or desktop), and create your own desktop app that will stay in your menubar.
Accessible provides an easy-to-access menu containing any of your favorite and most important files. This allows you to bypass Finder or any other file system app, and focus on what really matters: getting things done. AppGenome 1.4.2 – Create your own URL-based apps. April 4, 2017 AppGenome lets you set your own URL, browser agent type (mobile or desktop), and create your own desktop app that will stay in your menubar, so you can forget about all those “app for” apps.
Yuriy Georgiev - $1.23
|
|
Dropshare 5 1 5 esv. An epigenome consists of a record of the chemical changes to the DNA and histone proteins of an organism; these changes can be passed down to an organism's offspring via transgenerational stranded epigenetic inheritance. Changes to the epigenome can result in changes to the structure of chromatin and changes to the function of the genome.[1]
Epigenome
Appgenome 1 428
The epigenome is involved in regulating gene expression, development, tissue differentiation, and suppression of transposable elements. Unlike the underlying genome, which remains largely static within an individual, the epigenome can be dynamically altered by environmental conditions.
Cancer[edit]
Epigenetics is a currently active topic in cancer research. Human tumors undergo a major disruption of DNA methylation and histone modification patterns. The aberrant epigenetic landscape of the cancer cell is characterized by a global genomic hypomethylation, CpG island promoter hypermethylation of tumor suppressor genes, an altered histone code for critical genes and a global loss of monoacetylated and trimethylated histone H4.
Epigenome research projects[edit]
As a prelude to a potential Human Epigenome Project, the Human Epigenome Pilot Project aims to identify and catalogue Methylation Variable Positions (MVPs) in the human genome.[2] Advances in sequencing technology now allow for assaying genome-wide epigenomic states by multiple molecular methodologies.[3] Micro- and nanoscale devices have been constructed or proposed to investigate the epigenome.[4]
An international effort to assay reference epigenomes commenced in 2010 in the form of the International Human Epigenome Consortium (IHEC).[5][6][7][8] IHEC members aim to generate at least 1,000 reference (baseline) human epigenomes from different types of normal and disease-related human cell types.[9][10][11]
Roadmap epigenomics project[edit]
One goal of the NIH Roadmap Epigenomics Project is to generate human reference epigenomes from normal, healthy individuals across a large variety of cell lines, primary cells and primary tissues. Data produced by the project, which can be browsed and downloaded from the Human Epigenome Atlas, fall into five types that assay different aspects of the epigenome and outcomes of epigenomic states (such as gene expression):
- Histone Modifications – Chromatin Immunoprecipitation Sequencing (ChIP-Seq) identifies genome wide patterns of histone modifications using antibodies against the modifications.[12]
- DNA Methylation – Whole Genome Bisulfite-Seq, Reduced Representation Bisulfite-Seq (RRBS), Methylated DNA Immunoprecipitation Sequencing (MeDIP-Seq), and Methylation-sensitive Restriction Enzyme Sequencing (MRE-Seq) identify DNA methylation across portions of the genome at varying levels of resolution down to basepair level.[13]
- Chromatin Accessibility – DNase I hypersensitive sites Sequencing (DNase-Seq) uses the DNase I enzyme to find open or accessible regions in the genome.
- Gene Expression – RNA-Seq and expression arrays identify expression levels or protein coding genes.
- Small RNA Expression – smRNA-Seq identifies expression of small noncoding RNA, primarily miRNAs.
Reference epigenomes for healthy individuals will enable the second goal of the Roadmap Epigenomics Project, which is to examine epigenomic differences that occur in disease states such as Alzheimer's disease.
Appgenome 1 42
See also[edit]
References[edit]
Appgenome 1 45
- ^Bernstein, Bradley E.; Meissner, Alexander; Lander,Eric S. (February 2007). 'The Mammalian Epigenome'. Cell. 128 (4): 669–681. doi:10.1016/j.cell.2007.01.033. PMID17320505.
- ^'Human Epigenome Project'. Archived from the original on 2011-07-16. Retrieved 2011-06-29.
- ^Milosavljevic, Aleksandar (June 2011). 'Emerging patterns of epigenomic variation'. Trends in Genetics. 27 (6): 242–250. doi:10.1016/j.tig.2011.03.001. PMC3104125. PMID21507501.
- ^Aguilar, Carlos; Craighead, Harold (October 4, 2013). 'Micro- and nanoscale devices for the investigation of epigenetics and chromatin dynamics'. Nature Nanotechnology. 8 (10): 709–718. Bibcode:2013NatNa..8.709A. doi:10.1038/nnano.2013.195. PMC4072028. PMID24091454.
- ^'Time for the epigenome'. Nature. 463 (7281): 587. Feb 2010. Bibcode:2010Natur.463Q.587. doi:10.1038/463587a. PMID20130607.
- ^Abbott, A (2010). 'Project set to map marks on genome'. Nature. 463 (7281): 596–597. doi:10.1038/463596b. PMID20162836.
- ^Bae, JB (2013). 'Perspectives of international human epigenome consortium'. Genomics Inform. 11 (1): 7–14. doi:10.5808/GI.2013.11.1.7. PMC3630389. PMID23613677.
- ^'BioNews - Human Epigenome project launched'.
- ^'France: Human epigenome consortium takes first steps'Archived 2015-07-08 at the Wayback Machine. 5 March 2010.
- ^Eurice GmbH. 'About IHEC'.
- ^Kanai, Yae; Arai, Eri (2014). 'Multilayer-omics analyses of human cancers: Exploration of biomarkers and drug targets based on the activities of the International Human Epigenome Consortium'. Frontiers in Genetics. 5: 24. doi:10.3389/fgene.2014.00024. PMC3924033. PMID24592273.
- ^Zhu, J.; et al. (2013). 'Genome-wide chromatin state transitions associated with developmental and environmental cues'. Cell. 152 (3): 642–654. doi:10.1016/j.cell.2012.12.033. PMC3563935. PMID23333102.
- ^Harris, R Alan; Wang, Ting; Coarfa, Cristian; Nagarajan, Raman P; Hong, Chibo; Downey, Sara L; et al. (September 19, 2010). 'Comparison of sequencing-based methods to profile DNA methylation and identification of monoallelic epigenetic modifications'. Nature Biotechnology. 28 (10): 1097–1105. doi:10.1038/nbt.1682. PMC2955169. PMID20852635.
Appgenome 1 4 X 4

External links[edit]
Retrieved from 'https://en.wikipedia.org/w/index.php?title=Epigenome&oldid=963580779'
